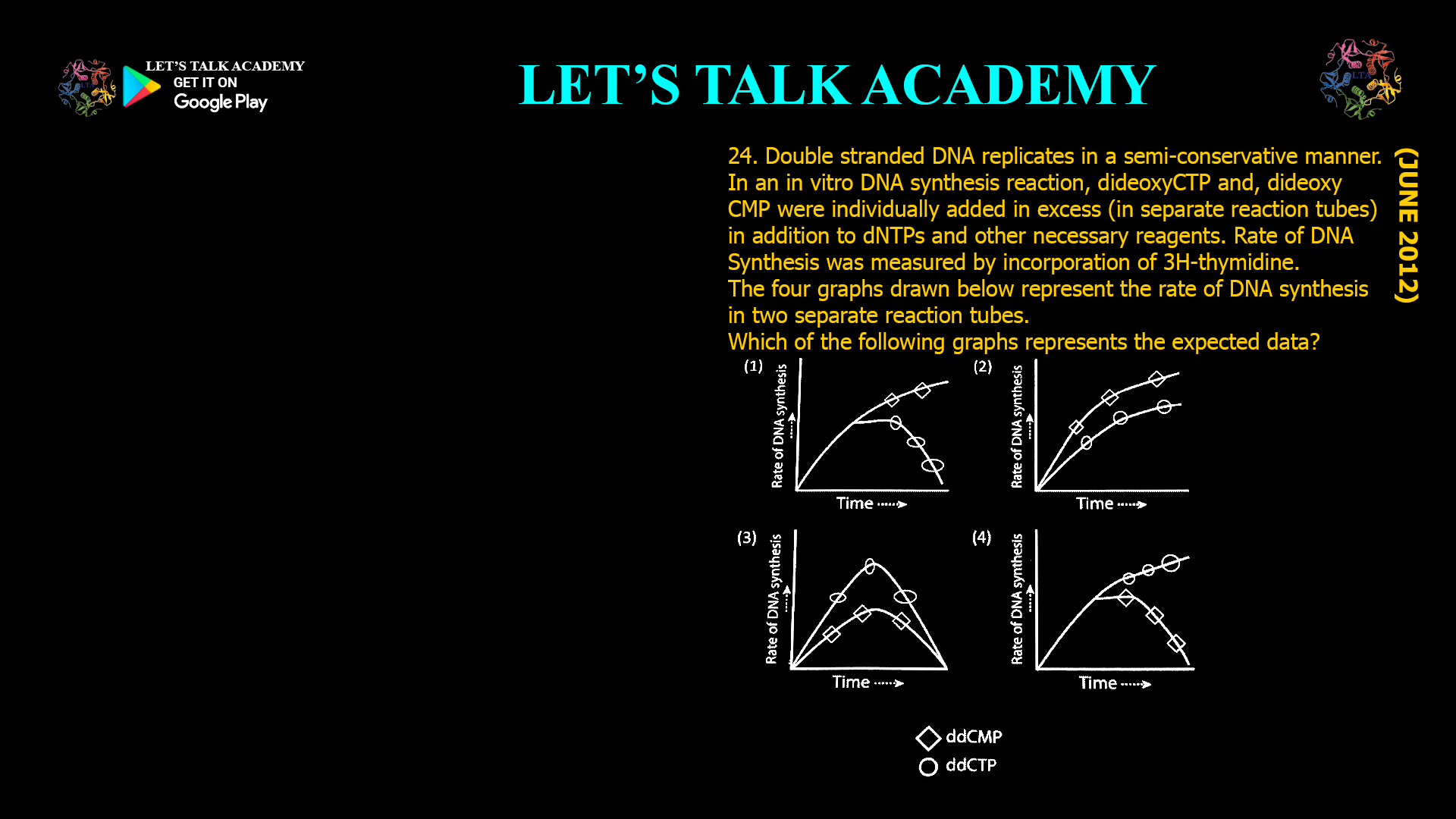
22. Match the chemical agents that interfere in oxidative phosphorylation process with their respective mode of action. Column I Column II (A) Antimycin A (i) Inhibits Fo component of ATP […]
Category: CSIR NET Life Science Previous Year Questions and Solution on DNA REPLICATION
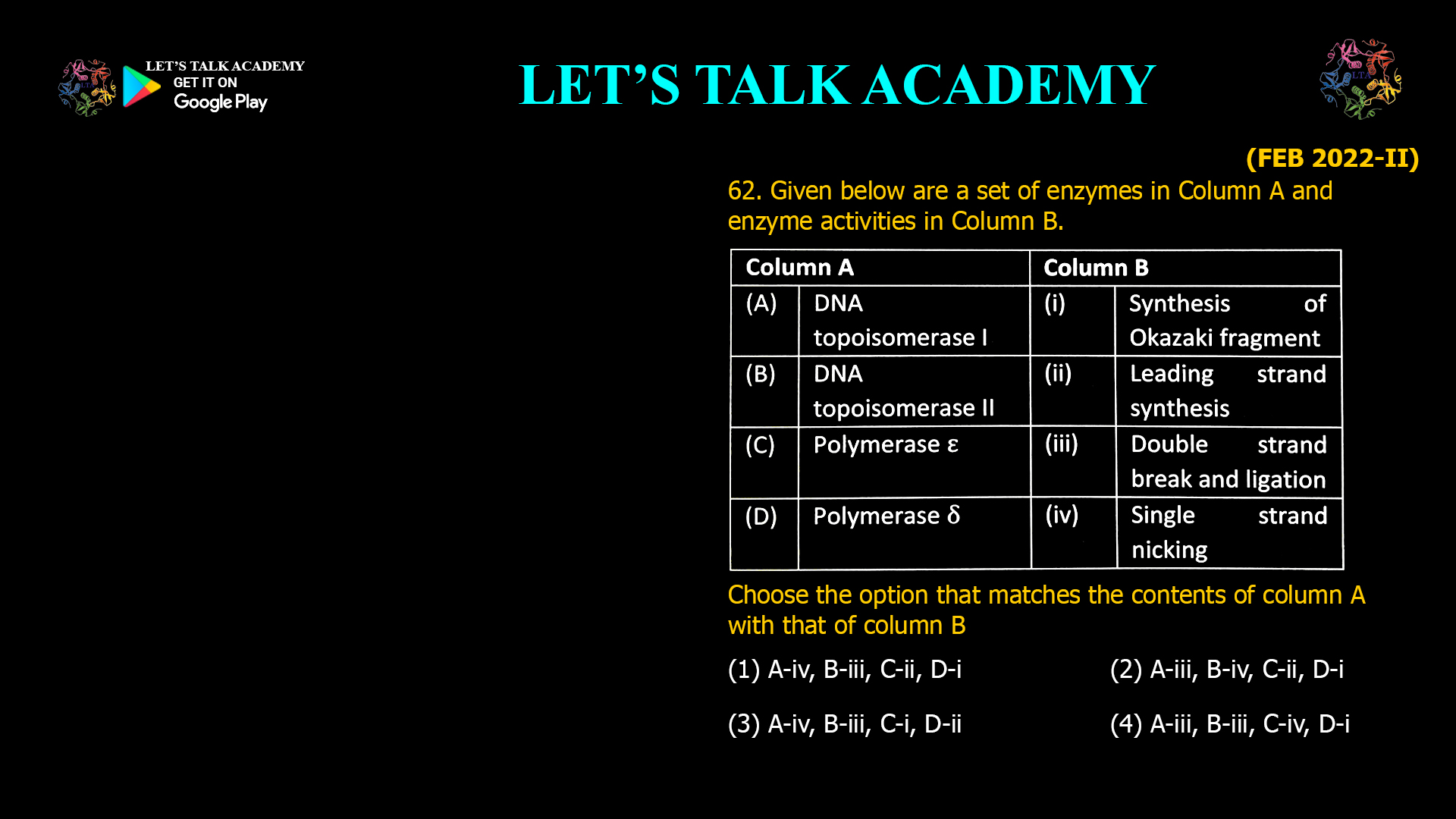
CSIR NET Life Science Previous Year Questions and Solution on DNA REPLICATION

Insights into Plasmid Curing and Segregation: Random Distribution vs Active Partitioning in Bacteria

Which DNA Replication Statements Are Incorrect? A Comparison of Prokaryotic and Eukaryotic Processes