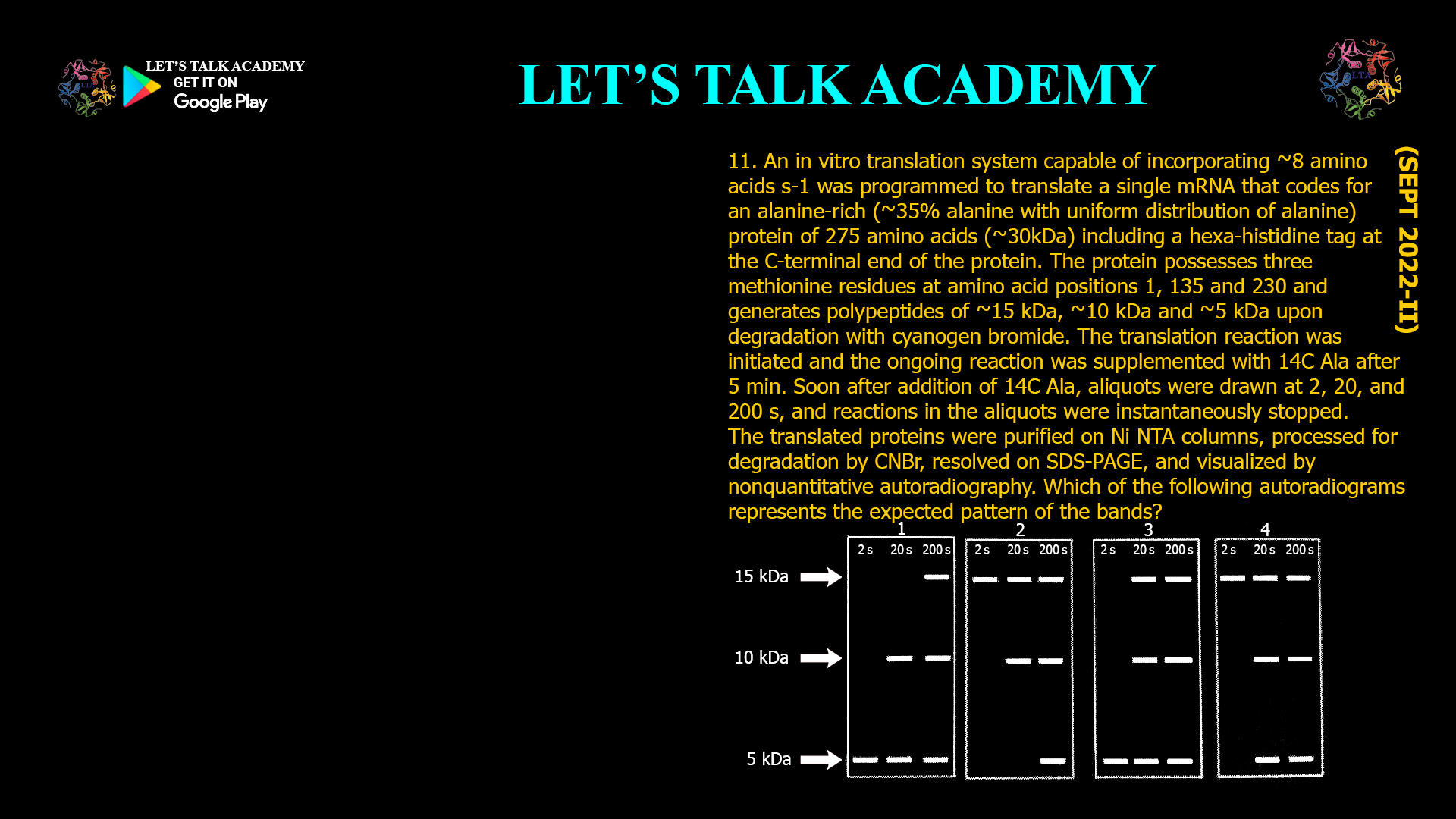
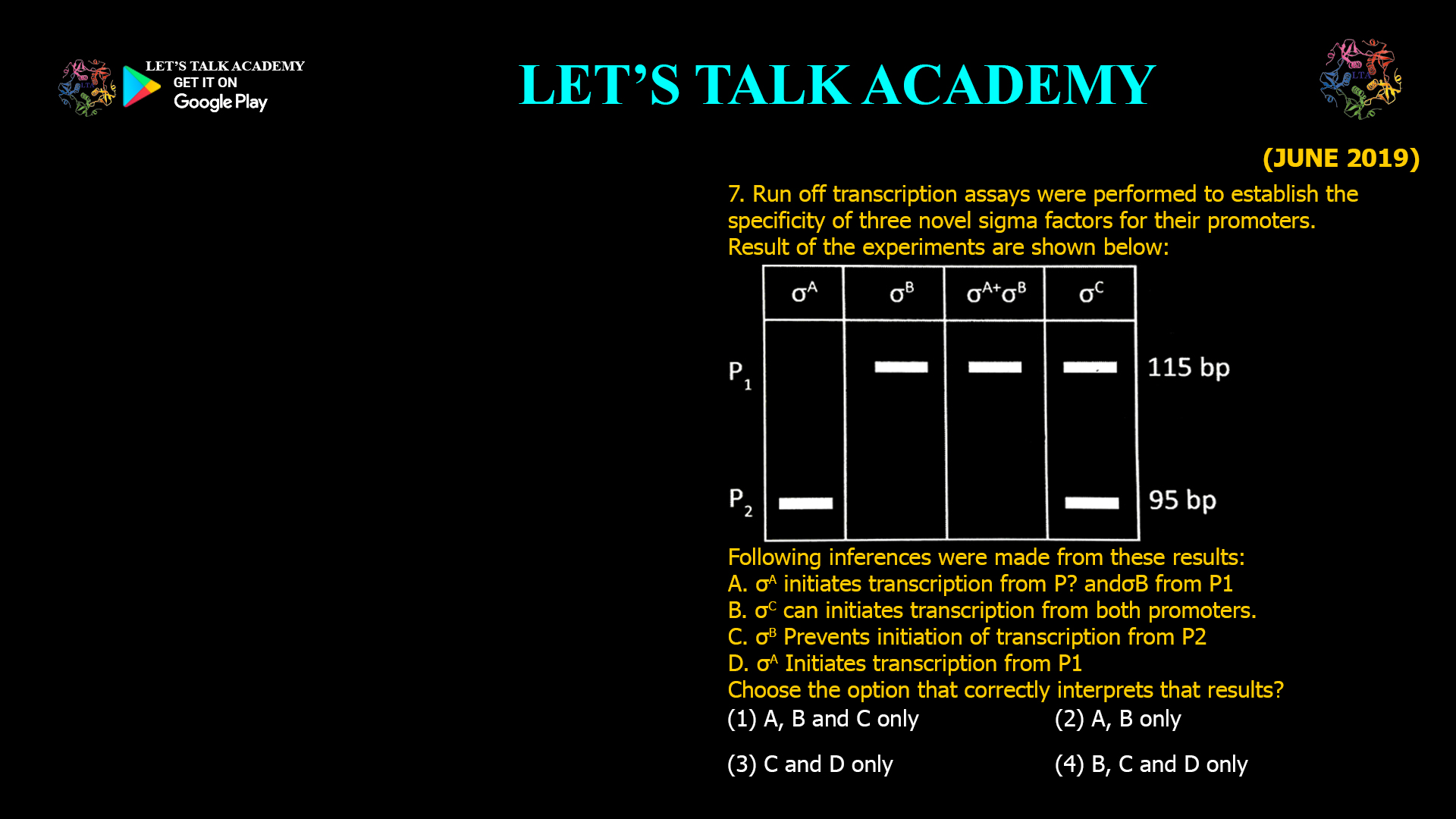
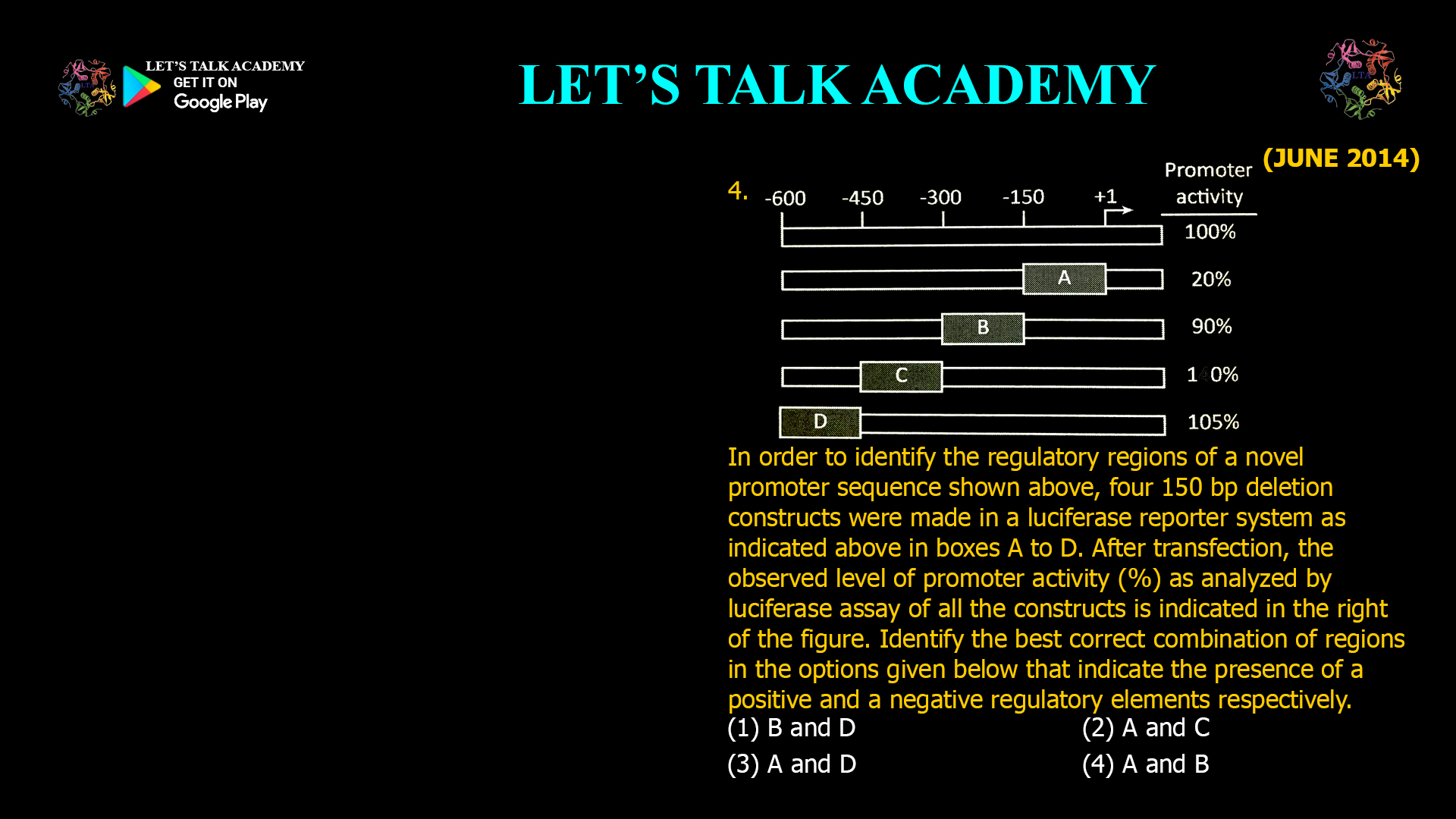
11. An in vitro translation system capable of incorporating ~8 amino acids was programmed to translate a single mRNA that codes for an alanine-rich (35% alanine with uniform distribution of […]
Category: CSIR NET Life Science Previous Year Questions and Solution on MOLECULAR TECHNIQUES
CSIR NET Life Science Previous Year Questions and Solution on MOLECULAR TECHNIQUES